Our latest article “Morphometric analysis of lungfish endocasts elucidates early dipnoan palaeoneurological evolution” was published yesterday in eLife.
In the way that many projects tend to go, the work for this article is several years in the making. I developed the seed of the idea during my time at Uppsala University (Sweden), and have been collaborating closely with Dr Tom Challands (University of Edinburgh) for several years to bring it to fruition. Together, Tom and I pooled our lungfish endocrania to more than double the number of those with “endocasts” known.
A cranial endocast is the internal space within the skull where the brain sits. As only the hard, bony parts of animals tend to fossilise we rely on the shape of these spaces to make inferences about the (now long gone) brain within.

Using synchrotron and micro-CT scanning technology, we were able to scan six new lungfish fossils to create virtual models of their endocasts. The fossils need to be exceptionally-preserved in 3D to be able to conduct these analyses. The fossils came from different sites from around the world (Australia, USA, Russia and Germany), but all hailed from the lungfish “heyday” of the Devonian Period (359-419 million years ago).
By combining these six new fossil models with all of the other lungfish endocasts known were were able to compile a dataset of 12 taxa which we then measured. The skulls of Devonian lungfish are highly variable in shape, and so we predicted that different endocast (brain) regions would undergo elongation as the external skulls changed shape.
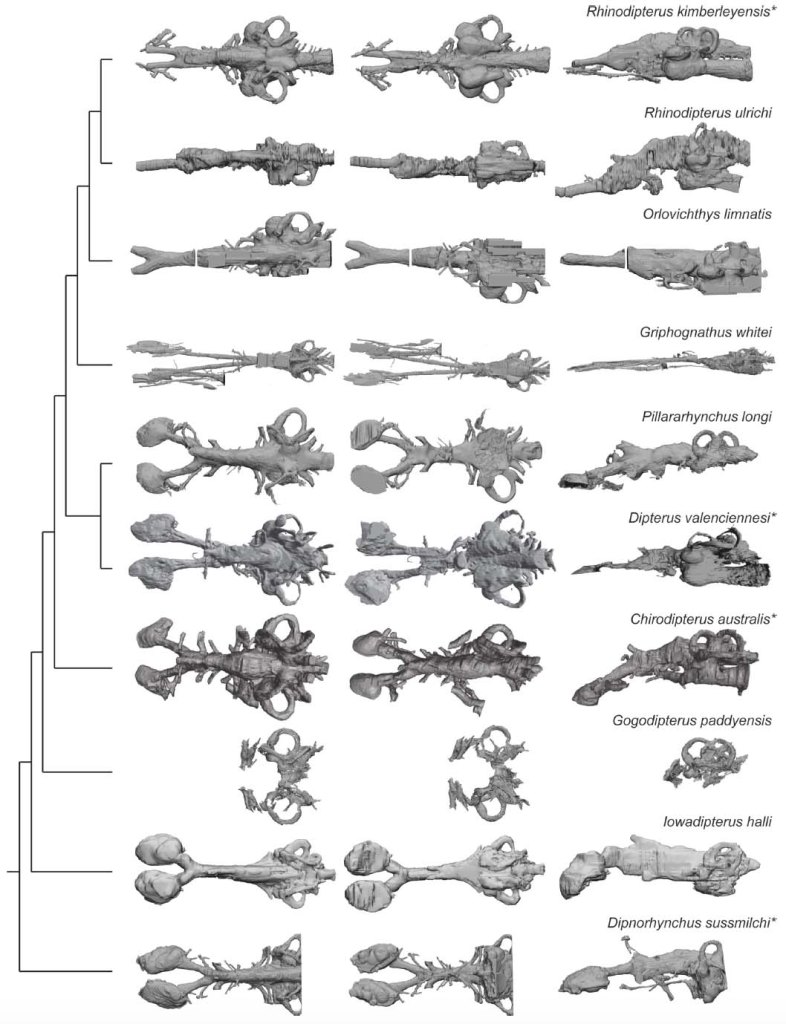
Along comes our heroic collaborator, Prof. Richard Cloutier (UQAR), who then had the unenviable task of making sense of our messy measurements and conducting some impressive morphometric analyses (a way of quantifying shape) despite our dataset having several missing variables (something not usually accommodated well in many morphometric analyses). We used several methods, including Bayesian Principal Component Analyses (BPCA) and PCA for incomplete data (InDaPCA) to untangle how the shapes of the endocasts differed from one another.
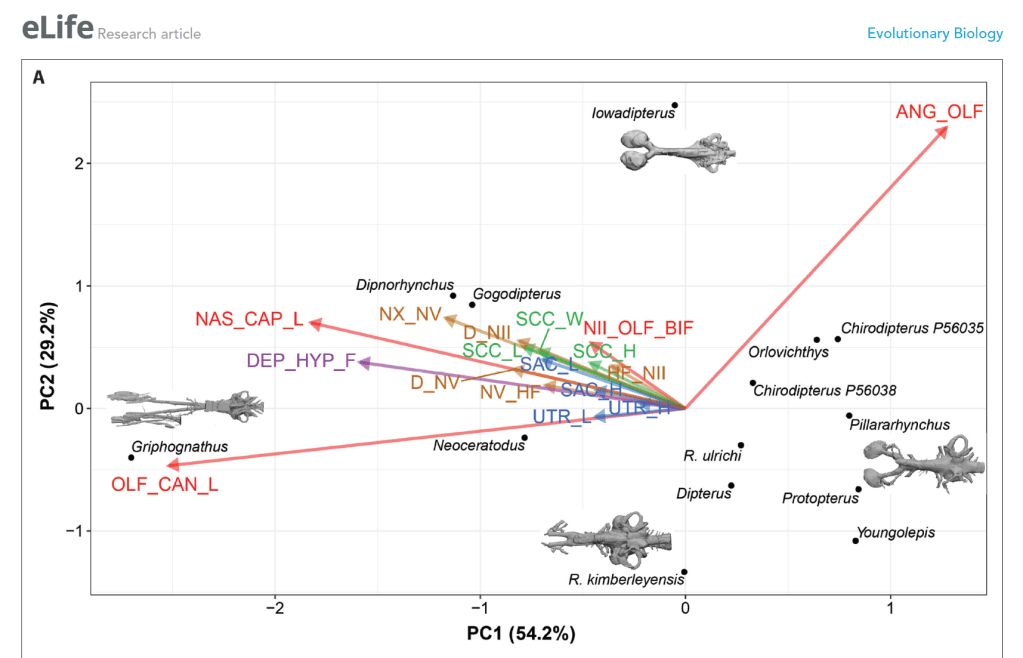
Our findings showed that contrary to our hypothesis where we thought different brain regions would tend to elongate within long skulls, most of the elongation (regardless of skull shape) tended to happen in the olfactory (sense of smell) region. We consider that sense of smell has remained an important sense throughout lungfish evolution.
We also uncovered some interesting things happening within the labyrinths (inner ears). The shape of inner ears can give a lot of information about an animal related to its sense of hearing, but also how it moves (locomotion). More investigation needs to be done but this may point to differing sensory requirements as lungfish evolved from deep sea animals to those living closer to shorelines in freshwater environments.
Ultimately we are continuing to learn more about brain evolution in this most fascinating group, the lungfish, which can in turn aid our interpretation of other groups -including our earliest fishy ancestors as they took the leap from water to land. Big thanks are due to all co-authors Tom Challands, Richard Cloutier, Laurent Houle, Per Ahlberg, Shaun Collin, John Long, as well as the editorial and reviewing team at eLife. If you would like to know more then I encourage you to read the article directly from eLife. All scan data is freely available for download from MorphoSource or Dryad.
